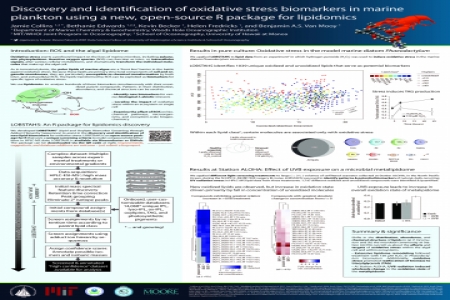
The lipids of marine plankton encompass a diversity of biochemical functions and taxonomic specificities that make them ideal molecular biomarkers in living biomass. While simple non-polar lipids such as free fatty acids (FFA) have formed the basis for many biomarker studies in fresh biomass, methods that enable the simultaneous profiling of both non-polar and intact polar lipids (IPL) have opened new avenues for characterization of environmental stressors. We demonstrate the application of a new, open-source lipidomics data analysis software package to identify a broad range of non-polar lipids, intact polar lipids, intact oxidized lipids (ox-IPL), and oxylipins associated with photooxidative stress in natural planktonic assemblages. Using the software, we identify lipid and oxylipin biomarkers associated with exposure of different marine microbial communities to (1) hydrogen peroxide (an artificial source of oxidative stress) and (2) ultraviolet radiation in the North Pacific Subtropical Gyre. The lipidomics software (LOBSTAHS; Lipid and Oxylipin Biomarker Screening Through Adduct Hierarchy Sequences) draws compound assignments from two customizable onboard databases that contain entries for more than 60,000 lipids, oxylipins and ox-IPL. The screening software and databases are integrated into a single R package which is freely available for download via GitHub or Bioconductor. LOBSTAHS is designed to be used with the existing metabolomics packages xcms and CAMERA